subrela.plot.matplotlib.draw_trace function¶
-
subrela.plot.matplotlib.draw_trace(ax, trace_data, orientation='vertical', kws=None)[source]¶ Draw trace lines.
- Parameters
ax (matplotlib.axes.Axes) – Subplot on which trace lines are drawn.
trace_data (pandas.DataFrame) – Data of trace lines returned by
subrela.plot.get_trace_datafunction.orientation ({'vertical', 'horizontal'}, optional) – Orientation of a tree. If
'vertical', the height direction of a tree corresponds to the y axis. If'horizontal', it corresponds to the x axis.kws (dict or None, optional) – Keyword arguments passed to
matplotlib.axes.Axes.plotmethod.
- Returns
lines (list[matplotlib.lines.Line2D]) – Trace lines.
Notes
If neither key
'color'nor key'c'is inkws,'k'is set to'color'. If neither key'linewidth'nor key'lw'is inkws,2is set to'linewidth'.Examples
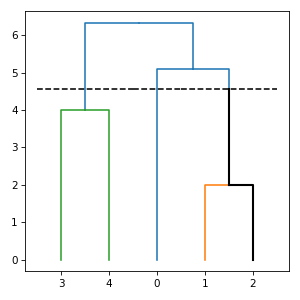
import numpy import matplotlib.pyplot from subrela.records import from_arrays from subrela.clustering import get_clusters from subrela.analysis import (get_strong_relevance_scores, get_weak_relevance_scores) from subrela.plot import get_dendrogram_data, get_trace_data from subrela.plot.matplotlib import draw_dendrogram, draw_trace scores = from_arrays([[False, False, False, True, True], [True, False, False, True, True], [False, True, False, True, True], [True, True, False, True, True], [False, False, True, True, True], [True, False, True, True, True], [False, True, True, True, True], [True, True, True, True, True]], [0.7, 0.7, 0.8, 0.8, 0.9, 0.9, 1., 1.]) X = numpy.array([[0, -5, -5, 6, 6], [0, -1, 1, -2, 2]]) Z = get_clusters(X) leaf_data, node_data, tree_data, cut_data = get_dendrogram_data( Z, groups=[0, 5, 6]) srs = get_strong_relevance_scores(scores, Z, clusters=[5]) wrs = get_weak_relevance_scores(scores, Z, 5) trace_data = get_trace_data(node_data, cut_data, wrs, tol=0.1) _, ax = matplotlib.pyplot.subplots(figsize=(4, 4)) draw_dendrogram(ax, leaf_data, tree_data, cut_data) draw_trace(ax, trace_data) matplotlib.pyplot.show()